Infection and Antimicrobial Resistance
Research in this theme is focused on improved prevention, detection and treatment of infection. Activities are concentrated on preventing the development of antimicrobial resistance through improved antimicrobial stewardship, improved detection of infection using molecular and sensor based technology, prevention of infection using novel anti-infective biomaterials and enhanced infection prevention and control strategies and improved treatment of infection through discovery of novel antibiotics and antibiotic adjuvants and markers for better evidence-based decisions on antibiotic selection.
Research Equipment Includes:
Illumina MiSeq
Next Generation Sequencing
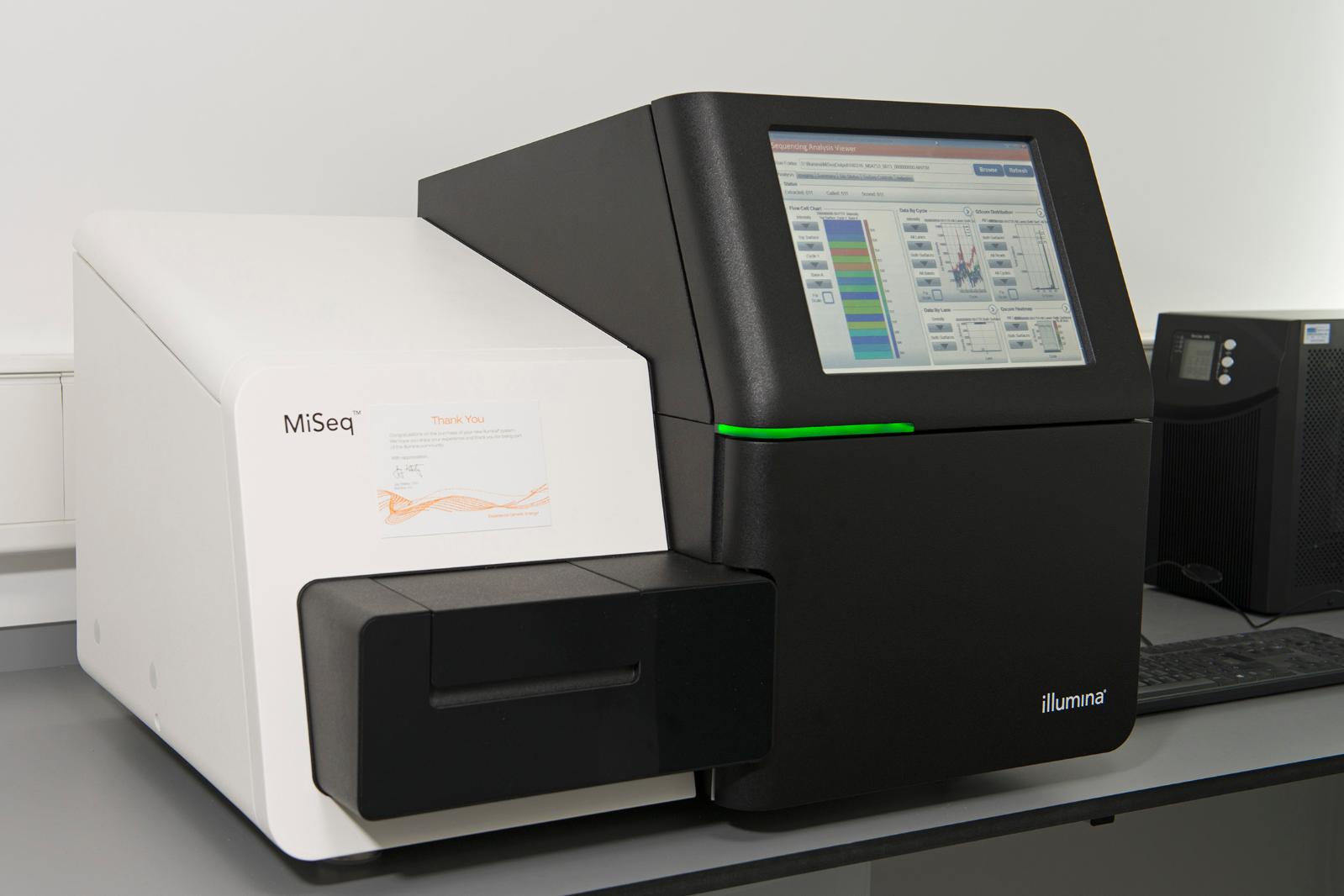

Integrates cluster generation, sequencing, and data analysis on a single instrument enabling the user to go from DNA to analyzed data in as little as 8 hours. Key applications consist of small genome sequencing, targeted gene sequencing and 16S metagenomic sequencing. Additional capabilities include; gene expression analysis with targeted RNA-Seq; targeted gene panels; De Novo sequencing; genotyping by sequencing; miRNA and small RNA analysis and DNA-protein interaction analysis with ChIP-Seq. Data upload feature to BaseSpace Sequence Hub enables cloud-based analysis and collaboration.
Located at MBC 03.014
Bookable only after suitable training
MagNA Pure 96 (Roche)
Automated Purifcation of Nucleic Acids
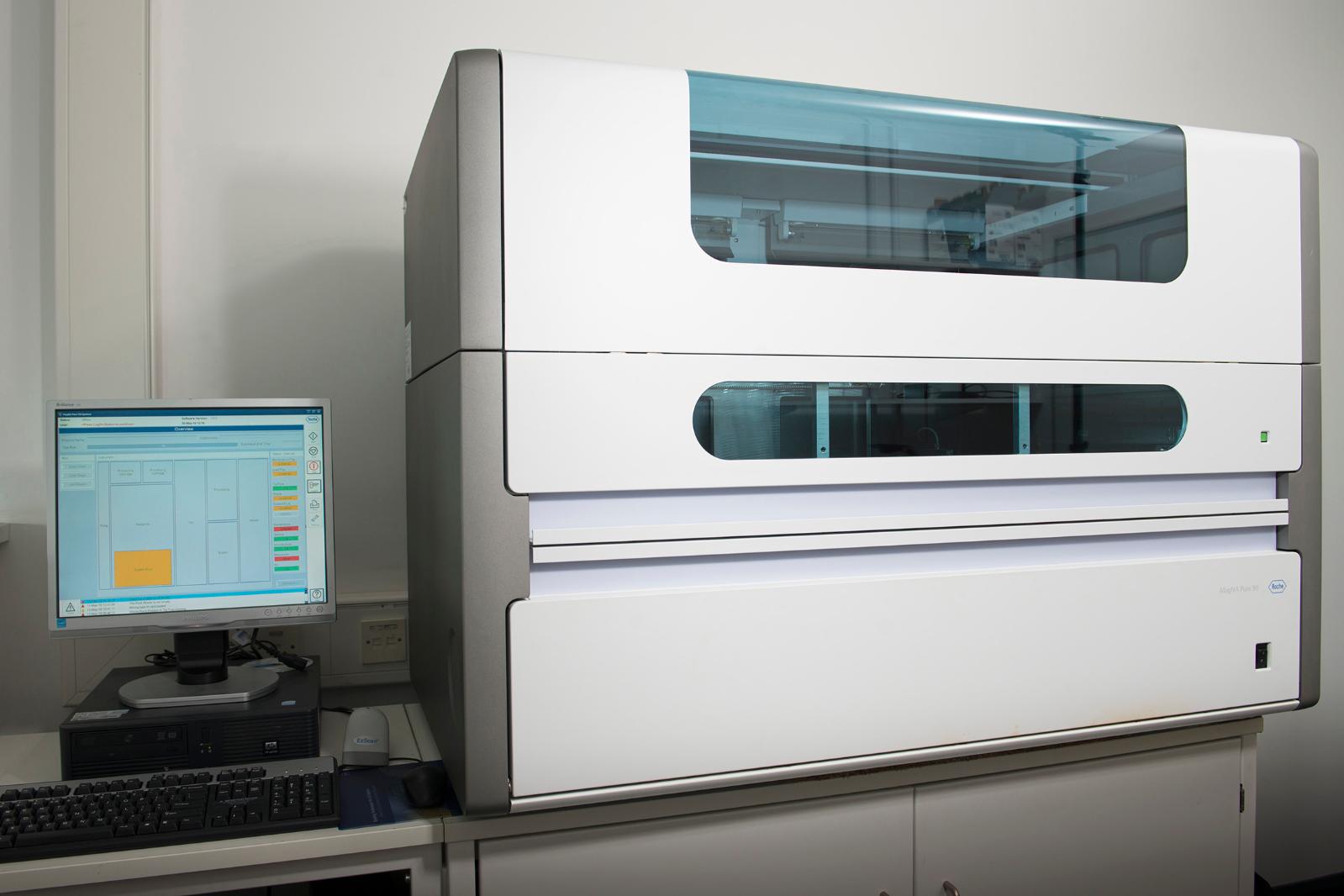

High-throughput purification of DNA, RNA, and viral nucleic acids from a wide range of starting materials using magnetic glass particle technology and prefilled, ready-to-use reagent cartridges. Up to 96 samples, ranging in volume from 50 to 500 µL, can be processed per run from a range of sample types such as whole blood, plasma, serum, urine, swabs, stool, sputum, BAL, CSF and bacterial cultures.
Located at MBC 03.005
Bookable only after suitable training
A34 and A45 Whitley Anaerobic Workstations
Anaerobic workstations for microbiological culture under anaerobic conditions
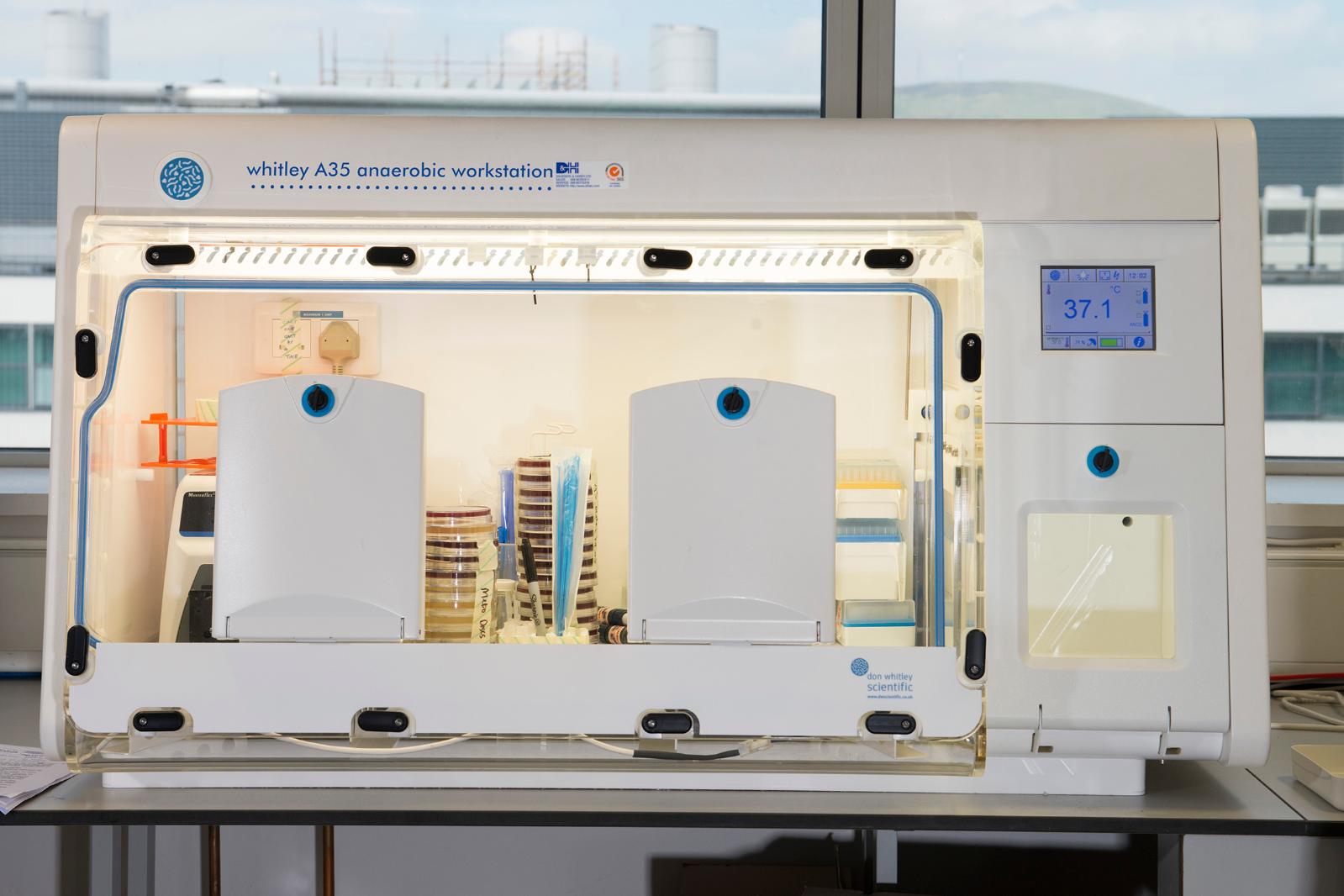
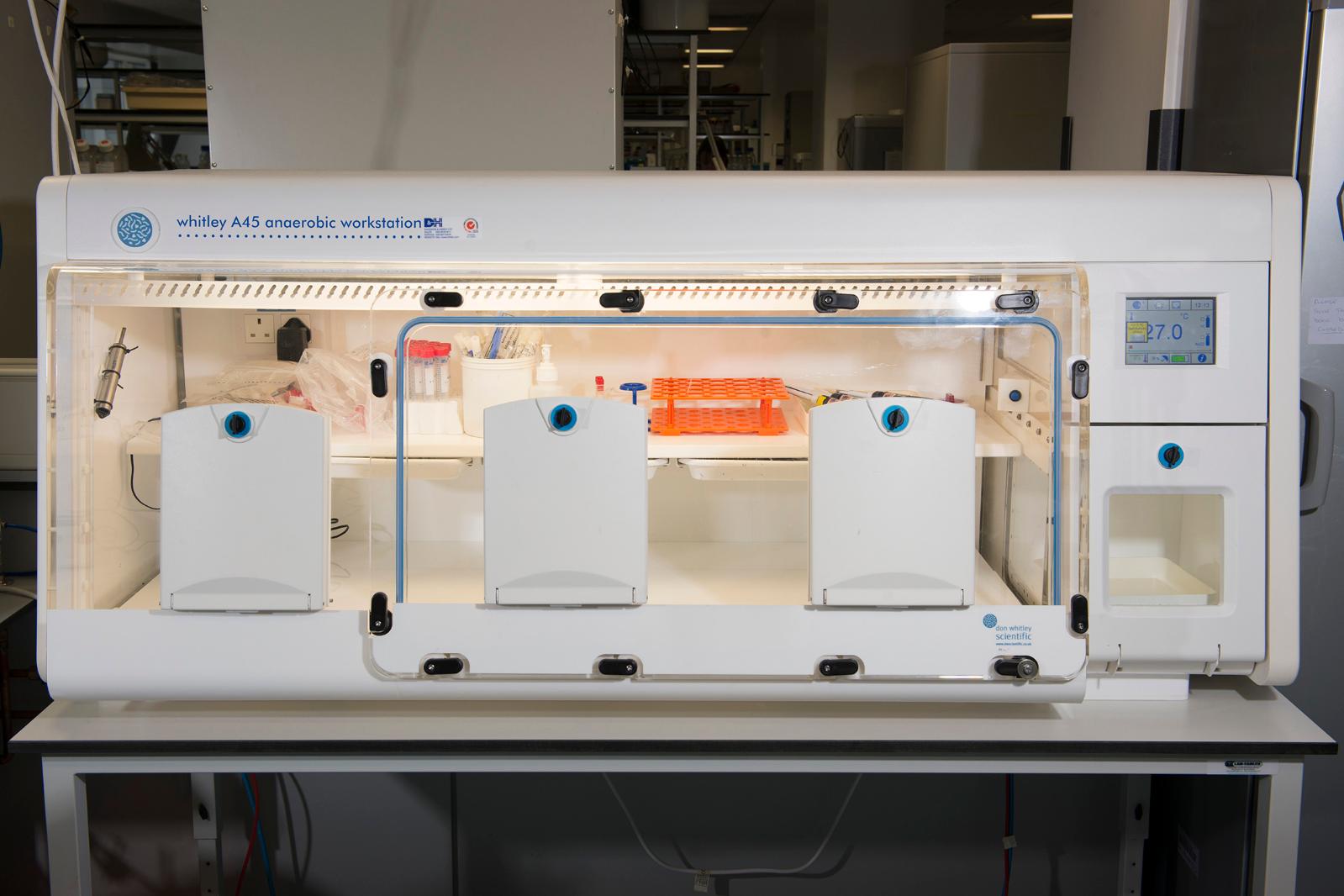


Processing of samples and incubation of bacterial strains without exposure to atmospheric oxygen, in a sustainable environment where parameters can be altered to create the required conditions. Instant access porthole system and built-in rapid airlock ensures samples can be transferred into the workstation quickly. Accommodates up to 600 x 90mm Petri dishes (A35) and over 750 x 90mm Petri dishes (A45). Removable front to facilitate the transfer of bulk samples and larger pieces of equipment into the workstation. Colour, touch-screen control panel for visual display of parameters such as temperature, humidity, and airlock cycle status.
Located at MBC 03.026
Bookable only after suitable training
LightCycler 480 II (Roche)
Real Time Polymerase Chain Reaction (RT-PCR)
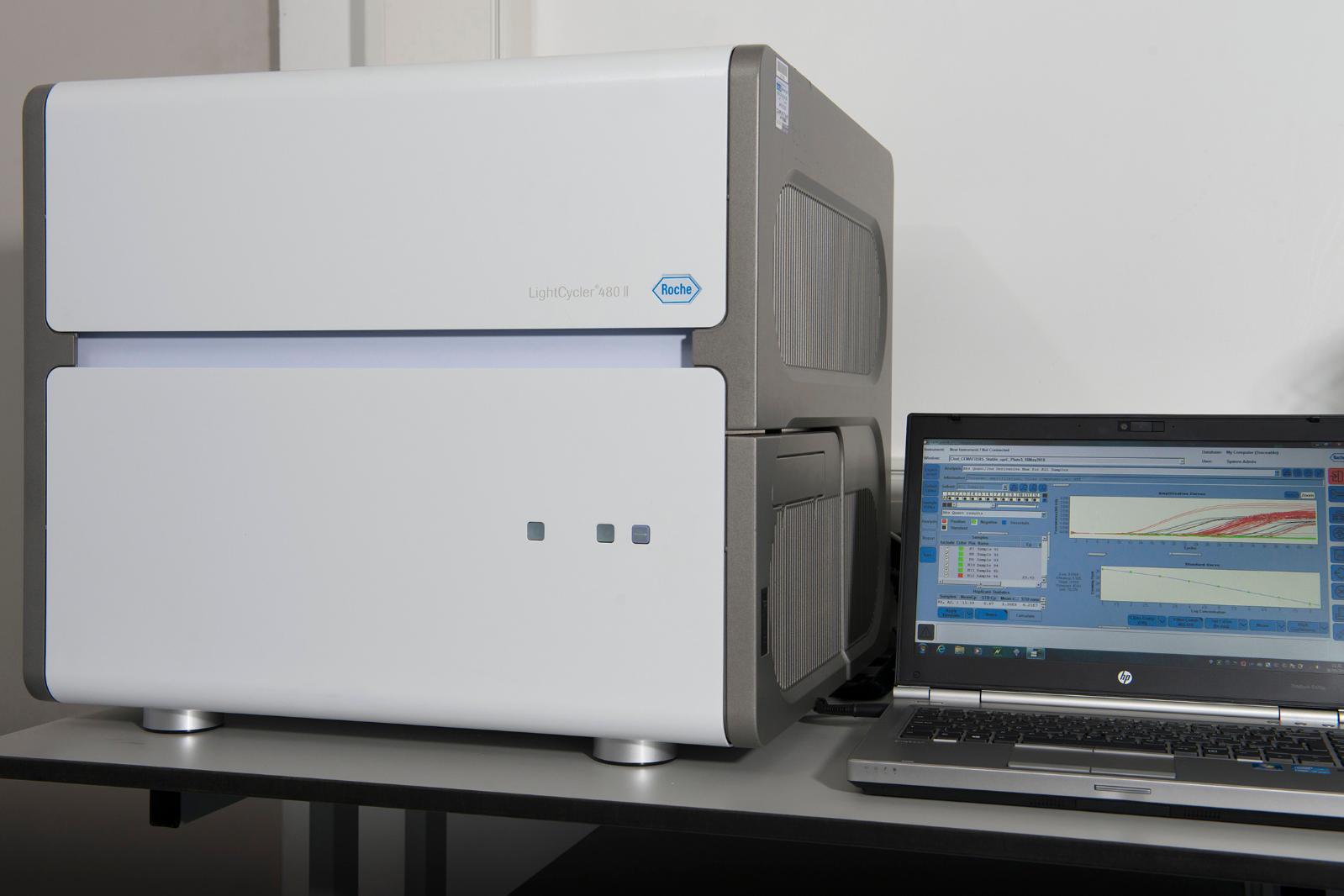

Qualitative and quantitative detection of nucleic acids, mutation scanning, and SNP analysis. Use basic and advanced methods for gene expression and genotyping based on endpoint analysis or melting curves. Probe based (e.g. TaqMan) and fluorescent dye based (e.g. Sybr Green) detection capabilities.
Located at MBC 03.026
Bookable only after suitable training
For more information on equipment and facilities in this research theme contact: Dr Andrew Lee, Research Technician, MBC 03012
E-mail: a.j.lee@qub.ac.uk
Tel: 02890 972091